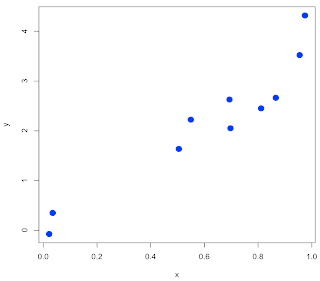
If we look at the errors, we see that they are dependent on the value of x! Naturally (b/c of mean=x[i]).
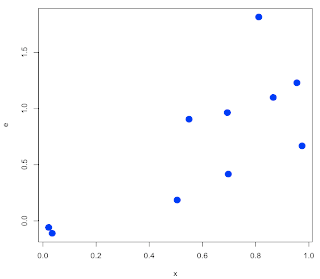
What I should have done is something like this:
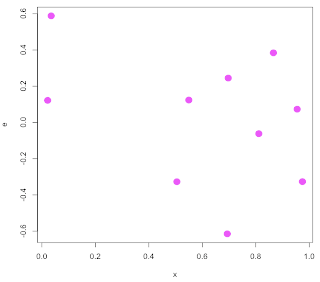
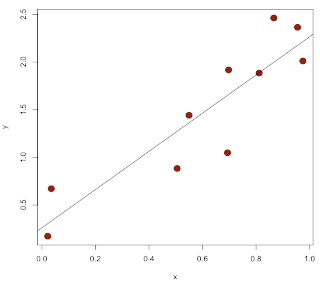
Nevertheless, I hope the essential points are clear:
• covariance is related to variance: cov(x,x) = var(x)
• correlation is the covariance of z-scores
• the slope of the regression line is: cov(x,y) / var(x)
• the regression line goes through x, y
• r ≈ cov(x,y) / sqrt(var(x)*var(y))
• the call to plot the line is abline(lm(y~x))
The proportionality for r is b/c we are not matching R's output for this. I think it is because we are missing a correction factor of n-1/n. I will have to look into that when I get back to my library at home.
With the change, we do a little better on guessing the slope: